Welcome to
Virtual Labs on Frontiers in Biochemistry
Copyright © 2017. All Rights Reserved. The Chinese University of Hong Kong.

Introduction
Enzymology is the study of enzymes, which has its roots in the early days of biochemistry. It is evolved from 19th century investigation of fermentation and digestion.
Research on fermentation is widely considered to have begun in the 18th century through determining that ethanol and CO2 are the principle products of sugar decomposed by yeast. The biological processes were caused by the action of chemical substances that were then known as "ferments" now the name "enzyme".
Virtual lab: Yeast Fermentation Experiment
Objective:
To find the ability of yeast in fermenting sugar in different conditions
To determine what is produced in fermentation
Principle:
Quantifying gas production is a standard approach to measure yeast fermentation. Although its approach is rather traditional, the apparatus is commonly used due to its low cost. Using this system, students can study effects of a wide range of parameters on the fermentation process.
Start the lab
Factors Affecting Enzyme Activity
Several factors affect the rate of enzymatic reaction - temperature, pH, enzyme concentration, substrate concentration, and the presence of inhibitors or activators.
Effect of Inhibitors on Enzyme Activity
Other factors, such as ionic strength, can also affect the enzymatic reaction. Each of these physical and chemical parameters must be considered and optimized in order for an enzymatic reaction experiment to be accurate and reproducible.
Effect of Inhibitors on Enzyme Activity
Enzyme inhibitors are substances which alter the catalytic action of the enzyme and consequently slow down, or in some cases, stop catalysis. There are three common types of enzyme inhibition - competitive, uncompetitive and mixed inhibition (or non-competition inhibition). Competitive inhibition involves inhibitor binding at an enzyme's substrate binding site. Uncompetitive inhibition involves inhibitor binding to enzyme-substrate complex. Mixed inhibition involves inhibitor binding to both the free enzyme and the enzyme-substrate complex.
Revision Questions
To access revision questions, please login CU learning system.
Course BCHE2000 Frontier in Biochemistry.
Topic: Biochemistry-history present and future\revision exercise.

Introduction
Proteins are the most abundant and functionally diverse molecules in living organisms.
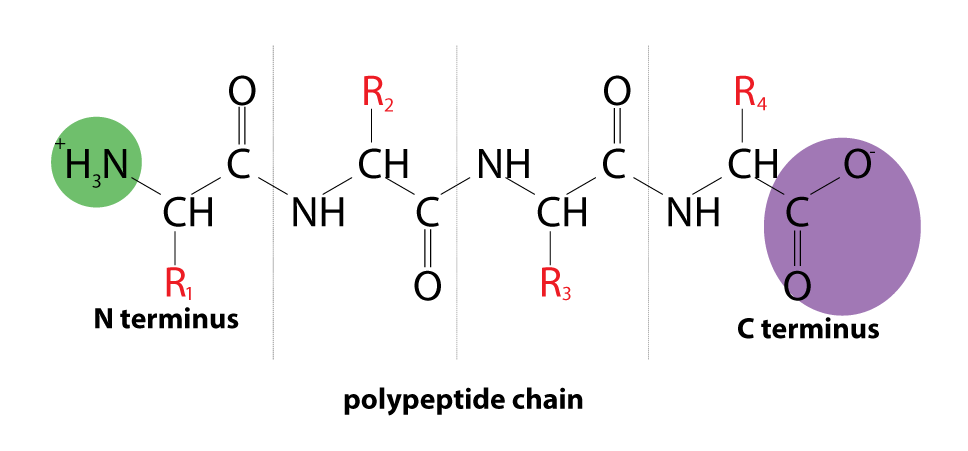
Proteins are macromolecules consisting of one or more polypeptides. Each polypeptide consists of a sequence of amino acids linked by peptide bonds. Polypeptide folds up to different structures with unique three dimensional conformation.
Proteins can be in different forms and possess different functions.
For examples:
- Enzymes - catalyze biochemical reactions. e.g. trypsin, subtilisin
- Transport proteins - e.g Hemoglobin
- Contractile or motile proteins - e.g. actin and myosin in skeletal muscle cells
- Structural proteins - to give strength and protection to biological structures. e.g. tendon, cartilage, leather, ligament, hair, fingernail, spider web ......
- Defense proteins - e.g antibodies
- Regulatory proteins - e.g. hormones, transcription factors
Amino acid sequence of a protein is rich in information. Mutation will result in malfunctioning of the protein or disease. Knowledge of a sequence provides valuable clues to its evolutionary relationship with other proteins, conformation and function.
Virtual lab: Protein sequencing
Objective:
To determine the protein sequence by Edman degradation method
Principle:
In 1950 Pehr Edman developed a method of protein sequencing. It involves sequential identification of amino acids from N to C terminal. Phenyl isothyocynate (PITC) reagent is used for the Edman degradation. The amino terminal of a protein can be identified by reacting the protein with PITC that forms a stable covalent linkage with the free amino group prior to hydrolysis with 6M HCl
Procedures:
Phenyl isothiocyanate (PITC, Edman reagent) reacts with the amino acid at the N-terminus of the peptide;
The PTH- amino acid is released from the peptide and identified by chromatography;
Cycles of labelling and release allow one to determine sequence of the peptide.
Start the lab
Mechanism of protein sequencing
Proteins Are Built from a Repertoire of 20 Amino Acids
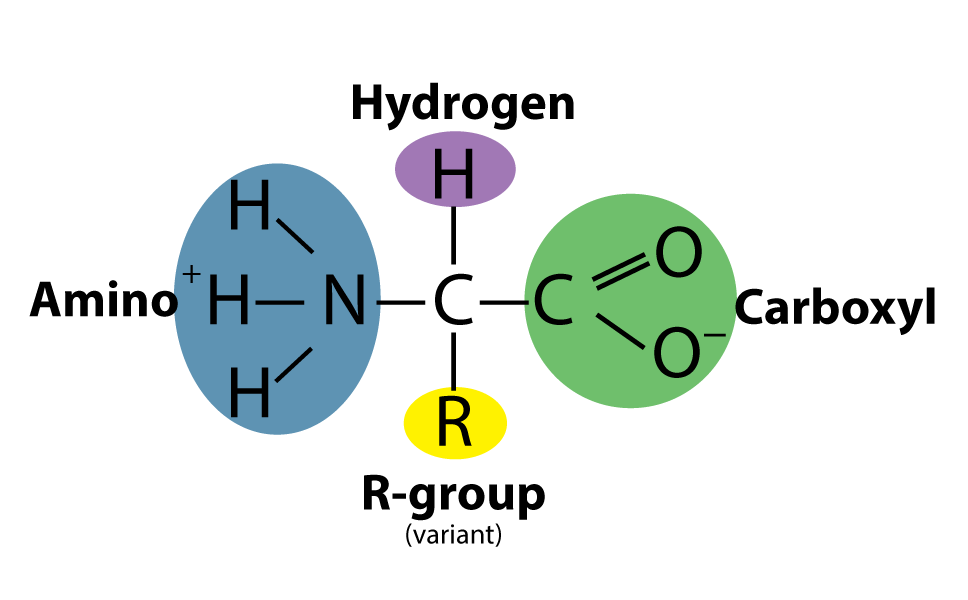
Amino acids are the building block of proteins. An amino acid consists of a central carbon atom, called α-carbon, linked to an amino group, a carboxylic acid group, a hydrogen atom, and a distinctive R group. The R group is often referred as the side chain.
Mechanism of protein sequencing
Amino Acid Sequence Can Be Determined by Automated Edman Degradation
The amino acid composition of a protein can be ascertained by hydrolyzing it into its constituent amino acids in 6 M HCl at 110℃. The amino acids can be separated by ion-exchange chromatography and quantitated by reacting them with ninhydrin or fluorescamine. Amino acid sequences can be determined by Edman degradation, which removes one amino acid at a time from the amino end of a peptide. Phenyl isothiocyanate reacts with the terminal amino group to form a phenylthiocarbamoyl derivative, which cyclizes under mildly acidic conditions to give a phenylthiohydantoin-amino acid and a peptide shortened by one residue. Automated repeated Edman degradation by a sequencer can analyze a peptide sequence of about 50 residues.
Longer polypeptide chains are broken into shorter ones for analysis by specifically cleaving them with a reagent such as cyanogen bromide, which splits peptide bonds on the carboxyl side of methionine residues.
Trypsin, which cleaves on the carboxyl side of lysine and arginine residues (if the next residue is not proline), is also useful in splitting proteins.
Revision Questions
To access revision questions, please login CU learning system.
Course BCHE2000 Frontier in Biochemistry.
Topic: Biochemistry-history present and future\revision exercise.
Introduction
Genetics is the study of genes, heredity, and genetic variation in living organisms. In the past century, many pioneering experiments have revealed the role of genes in heredity.
As life is basically a complex set of chemical reactions, so studying the way in which genes are relevant to these reactions is an important way of dissecting this complex chemistry.
Virtual lab: DNA sequencing
Objective:
To determine the DNA sequence by Sanger's method
Principle:
Sanger's sequencing method uses dideoxynucleotides (ddNTP) to interrupt DNA synthesis (also called chain-terminator method). Classically, a radioactive primer is used. In this experiment, the primers used in each of the four chain extension reactions are radioactively labelled. In each of four chain extension reactions, radioactively labelled primer is associated with a particular ddNTP. For example, the radioactively labelled primer in one chain reaction is terminated with ddCTP (dideoxycytidine triphosphate), while the radioactively labelled primer in another chain reaction is terminated with ddATP (dideoxyadenosine triphosphate) and so on. Each dideoxynucleotide (ddNTP) used gives all the fragments terminating in that particular nucleotide. The resulting DNA fragments are then separated by size in an electrophoresis gel. The DNA sequence can be read directly from an autoradiogram of the electrophoresis gel.
Start the lab
1. DNA synthesis by DNA polymerase
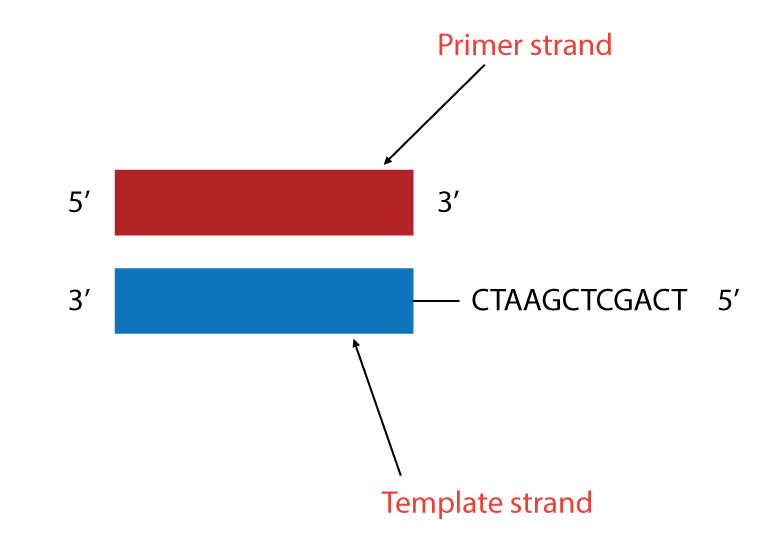
DNA sequencing by Sanger's method makes use of the mechanism of DNA synthesis by DNA polymerase. DNA polymerase requires both a primer strand (a short oligonucleotide strand), to which nucleotides are added, and a template strand to guide selection of each new nucleotide.
2. Interrupt DNA synthesis
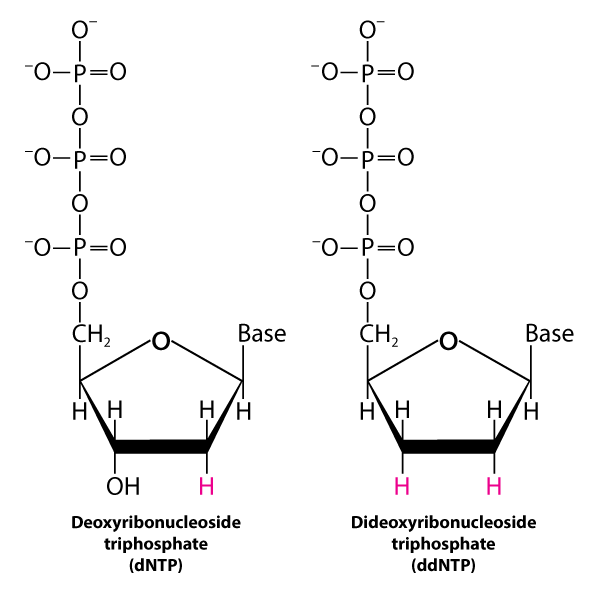
The Sanger's sequencing procedure uses dideoxynucleotide triphosphate (ddNTP) analogs to interrupt DNA synthesis. When a ddNTP is inserted in place of a dNTP, strand elongation is halted after the analog is added, because it lacks the 3-hydroxyl group needed for the next extension.
3. Initiation labelling
The DNA to be sequenced is used as the template strand, and a short primer, radioactively labeled, is annealed to it (i.e.: initiation labelling  ).
).
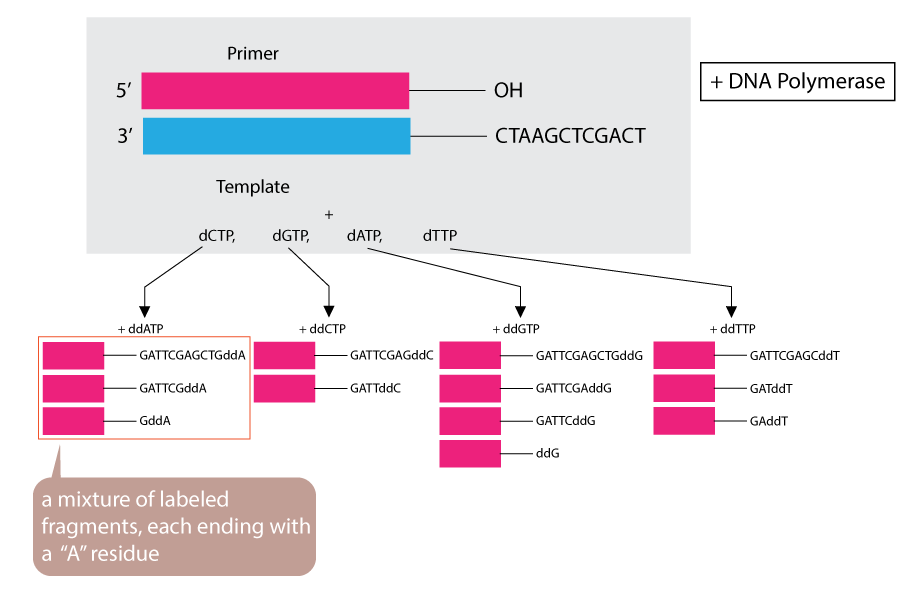
By addition of small amount of a single ddNTP, for example ddATP,to an otherwise normal reaction system, the synthesized strands will be prematurely terminated at some locations where dA normally occurs. Given the excess of dATP over ddATP, the chance of adding the analog to the growing chain is small.
(P.S.: For the radioactive labeling, there are two methods,
Initiation labelling: labelling the primer
Termination labelling: labelling the individual ddNTP)
4. Labelled fragment produced
However, ddATP is present in sufficient amount to ensure that each new strand has a high probability of acquiring one ddA at some point during synthesis. The result is a solution containing a mixture of labeled fragments, each ending with an “A” residue.
5. Electrophoresis
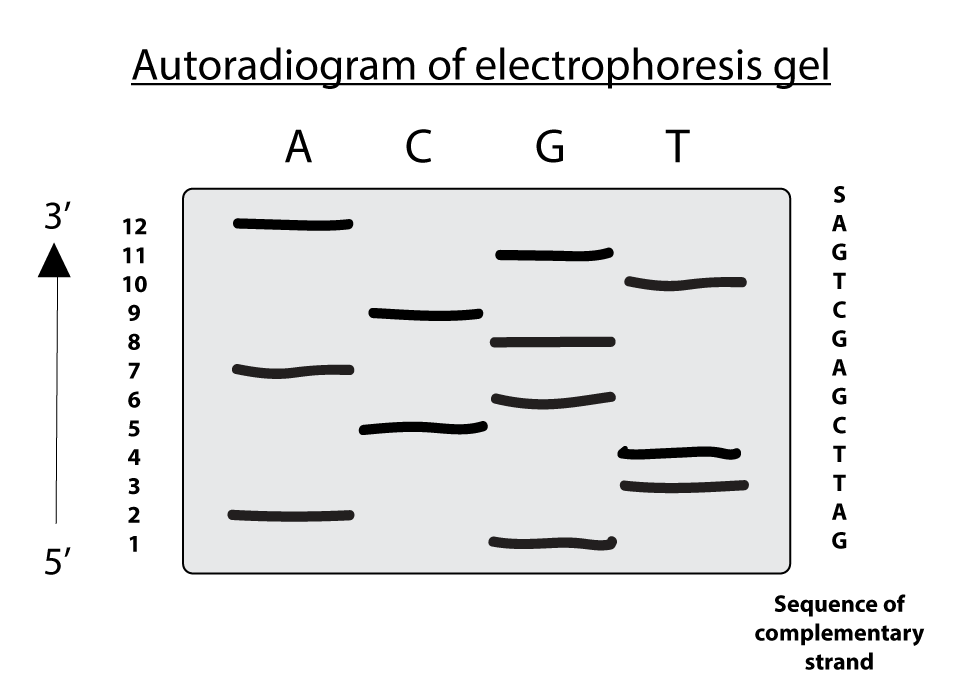
Each "A" residue in the sequence generates a set of fragments of a particular length, such that the different-sized fragments, separated by electrophoresis, reveal the location of "A" residues. This procedure is repeated separately for each of the four ddNTPs, and the sequence can be read directly from an autoradiogram of the electrophoresis gel. Because shorter DNA fragments migrate faster, the fragments near the bottom of the gel represent the nucleotide positions closest to the primer (the 5' end), and the sequence is read (in the 5' to 3' direction) from bottom to top. Note that the sequence obtained is the strand complementary to the strand being analyzed.
Revision Questions
To access revision questions, please login CU learning system.
Course BCHE2000 Frontier in Biochemistry.
Topic: Biochemistry-history present and future\revision exercise.

Introduction
Non-coding RNAs make up the majority (98%) of the transcriptome, and understanding the significance of this type of RNA can help us better understand nature and its application in our daily life.
Several different classes of regulatory RNAs with important functions have been discovered including small RNAs such as microRNAs and long noncoding RNAs (lncRNAs).
Classification of lncRNAs based on the sizes.

Long noncoding RNAs (lncRNAs) have gained widespread attention in recent years as a potentially new and crucial layer of biological regulation.
For example, X-inactive specific transcript (Xist) regulates allelic expression by X chromosome inactivation.
Virtual lab: X chromosome inactivation (Lyonization)
Objective:
To study X chromosome inactivation process in the embryotic development of tortoiseshell cats
Principle:
X chromosome inactivation takes place early in development at the blastocyst stage. The silencing of one chromosome at this stage is random, and each cell makes an independent choice of which chromosome to inactivate.
Once the "decision" is made, all descendants of that cell will keep the same pattern of inactivation in future cell division. The resulting mosaic-like pattern of expression can be seen in tortoiseshell cats.
Start the lab
Regulation of allelic expression
Mechanism of X chromosome inactivation
One of the studies on biological function for lncRNAs occurs in the epigenetic regulation of allelic expression is X chromosome inactivation (XCI), in which one of the two X chromosomes in females is heterochromatinized and silenced (the inactive X) such that only one X remains active and is expressed in each female cell (the active X).
Mechanism of X chromosome inactivation
Types of XCl
There are two types of XCI: 1) imprinted XCI in the fertilized embryo and extraembryonic tissues, where the paternal X is always inactivated; and 2) random XCI, occurring in the inner cell mass, where either the paternal or the maternal X is randomly chosen for inactivation, leading to a mosaic female. XCI in placental mammals is largely controlled by a lncRNA loci known as the X-inactivation center (Xic)
Initiation of XCI
The Xic contains several genetic elements essential for the transcription initiation of long noncoding RNAs that are involved in XCI. Initiation of XCI requires accumulation of a non-translated Xist RNA that coats the X chromosome. This is followed by various epigenetic changes on the future inactive X chromosome (Xi) that contributes to chromosome silencing.
Revision Questions
To access revision questions, please login CU learning system.
Course BCHE2000 Frontier in Biochemistry.
Topic: Non-coding RNA in development\revision exercise.